|
The one with the highest count is then assigned as the gene's primary function, the second-highest count gets assigned as the secondary function. For each gene we add up how many annotations fall under each high-level GO term. We assign each such GO term annotation to one of the high-level terms listed above. Using YeastMine we collect all GO term annotations for each protein-coding gene in SGD that trace back to one of the following "high-level" GO terms:Īll these terms fall under the GO term cellular process, and DNA replication falls under cellular metabolic process. GEMMER predicts functions for protein-coding genes using GO term annotations in SGD. Building the database Assignment of primary and secondary function through GO terms Excel and GEXF export formats may be imported for further analysis into Gephi and Cytoscape.Īll of the data GEMMER uses to visualize interaction networks is stored in an SQLite database.Data export for interaction networks to SVG, JSON, GEXF and Excel.Data visualization using interactive D3 (not 3D!) drawings.Interaction networks with localization, abundance and transcription timing information incorporated.Otwinowski: 'High-resolution timing of the cell-cycle regulated gene expression', PNAS 104, 16892 (2007). Specifically, we use the peak cell cycle phase transcription data from: M.SCEPTRANS (Saccharomyces Cerevisiae Periodic Transcription Server).YeastGFP (Yeast GFP Fusion Localization Database).CYCLoPs (Collection of Yeast Cells Localization Patterns).Effortless data integration from 3 databases. 
GEMMER aids (modeling) research by providing: After generating datasets and visualizations using GEMMER, the user may download the data and readily import it into for example Cytoscape for further analysis and ultimately model building. Furthermore, all this data may be inspected online and downloaded at the user's convienence.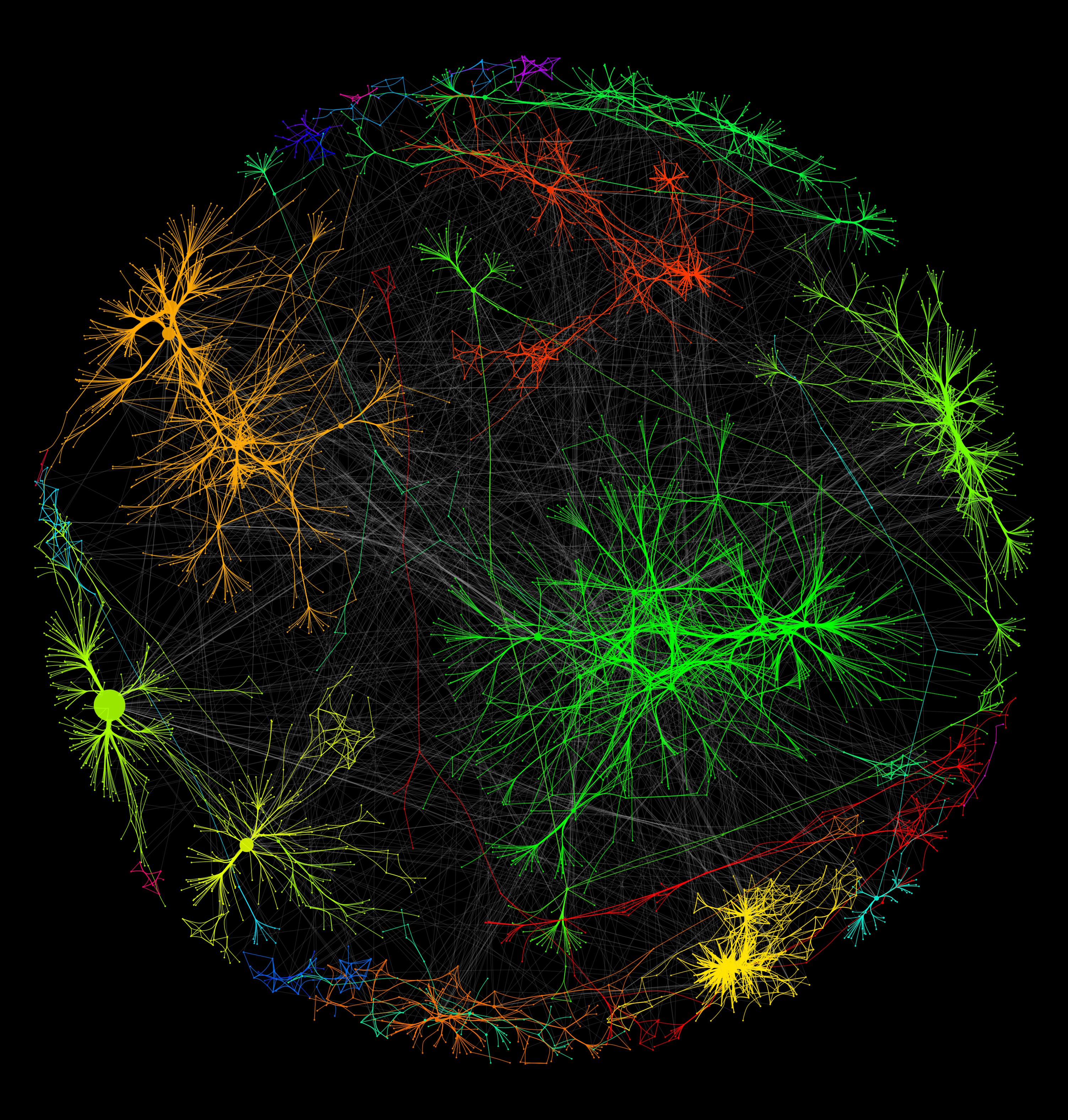  
The visualizations are genome-wide and multi-scale in the sense that the visualizations allow compartment localization, cell cycle transcription and functional data to be projected onto the network using data from various other databases. precious stones, which we thought apt to the process of discovering new biological insights which we hope GEMMER facilitates. In addition to being an acronym as described above, the word "gemmer" refers to mining of gems ( see the Merriam-webster definition), e.g. GEMMER ( GEnome-wide tool for Multi-scale Modeling data Extraction and Representation) aims to generate publication-quality visualizations of interactions between protein-coding genes in Saccharomyces cerevisiae and serve as a data-integration hub.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
